|
Biodiversity Information Science and Standards : Conference Abstract
|
|
Corresponding author: Lyubomir Penev (penev@pensoft.net)
Received: 30 Mar 2019 | Published: 13 Jun 2019
© 2019 Lyubomir Penev, Teodor Georgiev, Mariya Dimitrova, Yasen Mutafchiev, Pavel Stoev, Robert Mesibov
This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Citation: Penev L, Georgiev T, Dimitrova M, Mutafchiev Y, Stoev P, Mesibov R (2019) Data Auditing, Cleaning and Quality Assurance Workflows from the Experience of a Scholarly Publisher. Biodiversity Information Science and Standards 3: e35019. https://doi.org/10.3897/biss.3.35019
|
|
Abstract
Data publishing became an important task in the agenda of many scholarly publishers in the last decade, but far less attention has been paid to the actual reviewing and quality checking of the published data. Quality checks are often being delegated to the reviewers of the article narrative, many of whom may not be qualified to provide a professional data review. The talk presents the workflows developed and used by Pensoft journals to provide data auditing, cleaning and quality assurance. These are:
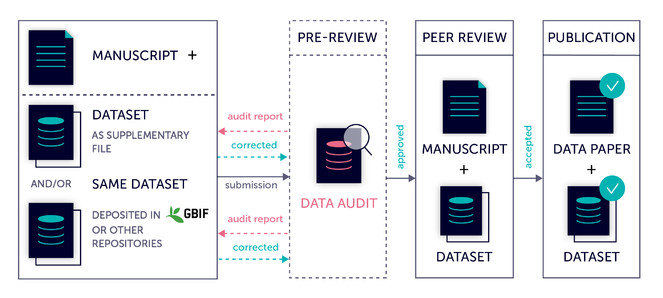
- Data auditing/cleaning workflow for datasets published as data papers (Fig.
1 , see also this blog). All datasets undergo an audit for compliance with a data quality checklist prior to peer-review. The author is provided with an audit report and is asked to correct the data flaws and consider other recommendations in the report. This check is conducted regardless of whether the datasets are provided as supplementary material within the data paper manuscript or linked from the Global Biodiversity Information Facility (GBIF) or another repository. The manuscript is not forwarded to peer review until the author corrects the data associated with it. This workflow is applied in all journals of the publisher’s portfolio, including Biodiversity Data Journal, ZooKeys, PhytoKeys, MycoKeys and others. - Automated check and validation of data within the article narratiive in the Biodiversity Data Journal provided during the authoring process in the ARPHA Writing Tool (AWT), and consequently, during the peer review process in the journal. Among others, the automated validation tool checks for compliance with the biological Codes (for example, a new species description cannot be submitted without designation of a holotype and the respective specimen record).
- Check for consistency and validation of the full-text JATS XML against the TaxPub XML schema. This quality check ensures a succesfull full-text submission and display on PubMedCentral, extraction of taxon treatments and their visualisation on Plazi's TreatmentBank and GBIF, indexing in various data aggregators, and so on.
- Human-provided quality check of the mass automated extraction of taxon treatments from legacy literature via the GoldenGate-Imagine workflow developed by Plazi and implemented for the purposes of the Arcadia project in a collaboration with Pensoft.
- Putting high-quality data in valid XML formats (Pensoft's JATS and Plazi's TaxonX) into a machine readable semantic format (RDF) to guarantee efficient extraction and transformation. Data from Pensoft's journals and Plazi's treatments are uploaded as semantic triples into the OpenBiodiv Biodiversity Knowledge Graph where it is modeled according to the OpenBiodiv-O ontology (
Senderov et al. 2018 ). Semantic technologies facilitate the mapping of scientific names in OpenBiodiv to GBIF's taxonomic backbone and the addressing of complex biodiversity questions.
Data auditing and cleaning workflow implemented in Pensoft for all datasets published as data papers.
We have realised in the course of many years experience in data publishing that data quality checking and assurance testing requires specific knowledge and competencies, which also vary between the various methods of data handling and management, such as relational databases, semantic XML tagging, Linked Open Data, and others. This process cannot be trusted to peer reviewers only and requires the participation of dedicated data scientists and information specialists in the routine publishing process. This is the only way to make the published biodiversity data, such as taxon descriptions, occurrence records, biological observations and specimen characteristics, truly FAIR (Findable, Accessible, Interoperable, Reusable), so that they can be merged, reformatted and incorporated into novel and visionary projects, regardless of whether they are accessed by a human researcher or a data-mining process.
Presenting author
Teodor Georgiev
References
-
OpenBiodiv-O: ontology of the OpenBiodiv knowledge management system.Journal of Biomedical Semantics9:5. https://doi.org/10.1186/s13326-017-0174-5